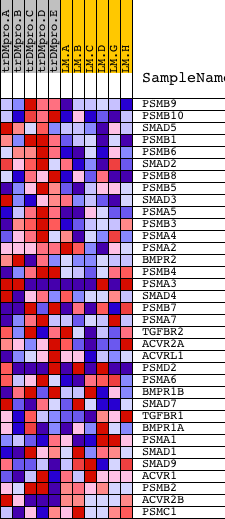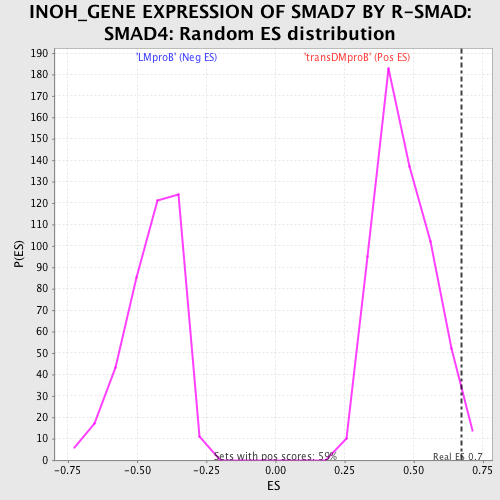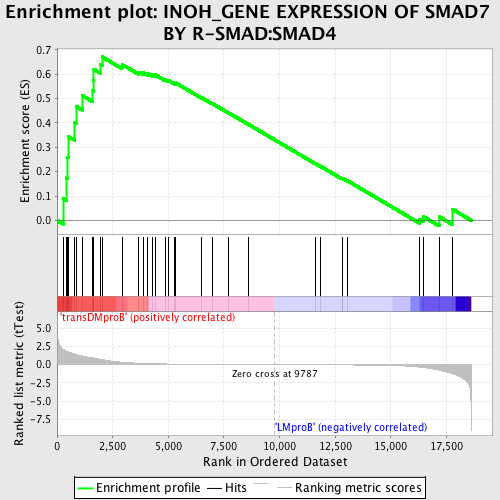
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_LMproB.phenotype_transDMproB_versus_LMproB.cls #transDMproB_versus_LMproB |
| Phenotype | phenotype_transDMproB_versus_LMproB.cls#transDMproB_versus_LMproB |
| Upregulated in class | transDMproB |
| GeneSet | INOH_GENE EXPRESSION OF SMAD7 BY R-SMAD:SMAD4 |
| Enrichment Score (ES) | 0.670851 |
| Normalized Enrichment Score (NES) | 1.4435588 |
| Nominal p-value | 0.02698145 |
| FDR q-value | 0.6913281 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PSMB9 | 23021 | 277 | 2.071 | 0.0893 | Yes | ||
| 2 | PSMB10 | 5299 18761 | 416 | 1.833 | 0.1742 | Yes | ||
| 3 | SMAD5 | 21621 | 469 | 1.756 | 0.2598 | Yes | ||
| 4 | PSMB1 | 23118 | 508 | 1.706 | 0.3436 | Yes | ||
| 5 | PSMB6 | 9634 | 775 | 1.454 | 0.4025 | Yes | ||
| 6 | SMAD2 | 23511 | 864 | 1.376 | 0.4671 | Yes | ||
| 7 | PSMB8 | 5000 | 1132 | 1.179 | 0.5120 | Yes | ||
| 8 | PSMB5 | 9633 | 1607 | 0.915 | 0.5326 | Yes | ||
| 9 | SMAD3 | 19084 | 1649 | 0.891 | 0.5753 | Yes | ||
| 10 | PSMA5 | 6464 | 1654 | 0.889 | 0.6198 | Yes | ||
| 11 | PSMB3 | 11180 | 1944 | 0.724 | 0.6407 | Yes | ||
| 12 | PSMA4 | 11179 | 2022 | 0.681 | 0.6709 | Yes | ||
| 13 | PSMA2 | 9631 | 2917 | 0.334 | 0.6396 | No | ||
| 14 | BMPR2 | 14241 4081 | 3678 | 0.190 | 0.6082 | No | ||
| 15 | PSMB4 | 15252 | 3866 | 0.166 | 0.6065 | No | ||
| 16 | PSMA3 | 9632 5298 | 4081 | 0.144 | 0.6023 | No | ||
| 17 | SMAD4 | 5058 | 4275 | 0.126 | 0.5982 | No | ||
| 18 | PSMB7 | 2795 14598 | 4408 | 0.117 | 0.5970 | No | ||
| 19 | PSMA7 | 14318 | 4864 | 0.092 | 0.5771 | No | ||
| 20 | TGFBR2 | 2994 3001 10167 | 5015 | 0.085 | 0.5733 | No | ||
| 21 | ACVR2A | 8542 | 5278 | 0.075 | 0.5630 | No | ||
| 22 | ACVRL1 | 22354 | 5322 | 0.074 | 0.5644 | No | ||
| 23 | PSMD2 | 10137 5724 | 6511 | 0.043 | 0.5026 | No | ||
| 24 | PSMA6 | 21270 | 6976 | 0.035 | 0.4794 | No | ||
| 25 | BMPR1B | 8661 15152 4451 | 7723 | 0.023 | 0.4404 | No | ||
| 26 | SMAD7 | 23513 | 8609 | 0.013 | 0.3934 | No | ||
| 27 | TGFBR1 | 16213 10165 5745 | 11615 | -0.020 | 0.2328 | No | ||
| 28 | BMPR1A | 8660 4450 | 11852 | -0.023 | 0.2212 | No | ||
| 29 | PSMA1 | 1627 17669 | 12805 | -0.037 | 0.1719 | No | ||
| 30 | SMAD1 | 18831 922 | 12826 | -0.038 | 0.1727 | No | ||
| 31 | SMAD9 | 15591 | 13032 | -0.042 | 0.1637 | No | ||
| 32 | ACVR1 | 4334 | 16280 | -0.327 | 0.0055 | No | ||
| 33 | PSMB2 | 2324 16078 | 16450 | -0.389 | 0.0160 | No | ||
| 34 | ACVR2B | 19276 | 17171 | -0.758 | 0.0154 | No | ||
| 35 | PSMC1 | 9635 | 17755 | -1.238 | 0.0463 | No |